Evolutionary dynamics of mitochondrial male sterility
|
Fabulous Collaborators: Lila Fishman, Findley Finseth, Jeff Mower, John Willis, Camille Barr, Eric Floro
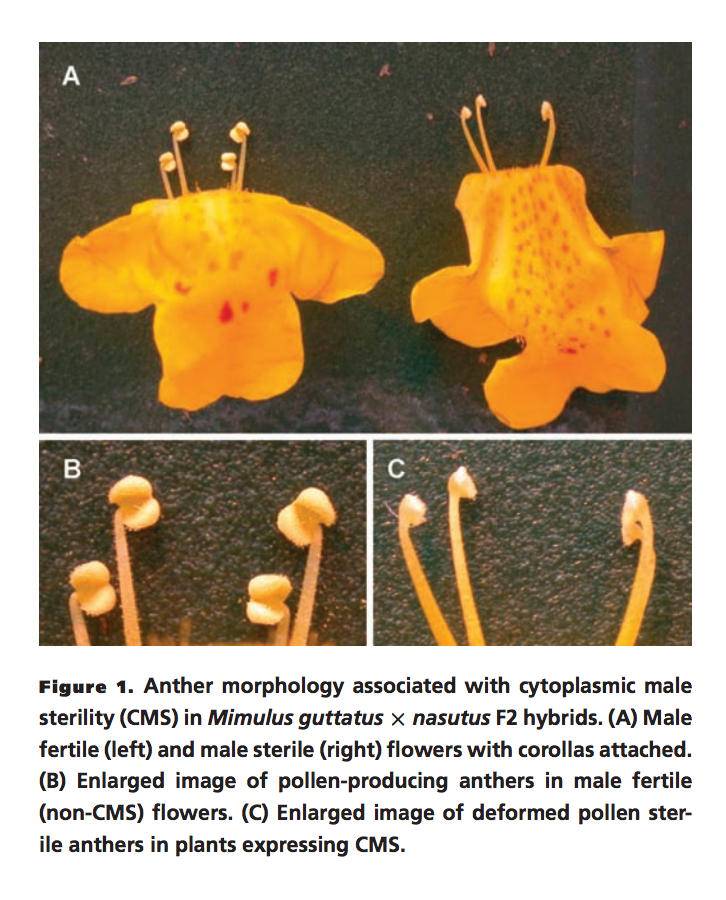
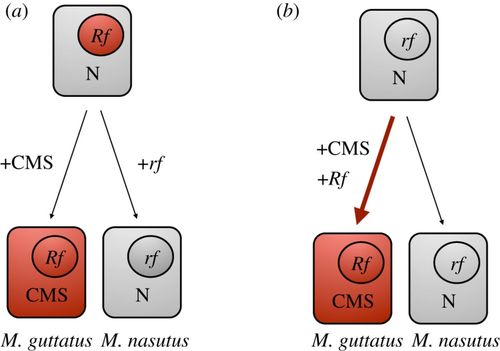
In eukaryotes, organellar genomes are typically uniparentally inherited, creating opportunity for intergenomic conflict with biparentally inherited nuclear genomes, despite the need for cooperation between these genomic components. For example, from the perspective of a maternally inherited mitochondrial genome, males are essentially expendable, and mutations that increase female but decrease male fertility spread readily within populations. This 'selfish' organellar evolution manifests in plants as mitochondrial male sterility (a.k.a. "CMS"), where carriers of expressed CMS genes are phenotypically female. Nuclear alleles have evolved to counteract CMS genes, restoring male function (and a hermaphroditic phenotype) to their carriers, but much is still unknown about the origins, dynamics and evolutionary consequences of this cyto-nuclear conflict over male fertility. It has potentially far-reaching consequences for organismal and genomic diversity, and may have profound effects on individual fitness, population differentiation, and even speciation. Our work on this has focused on populations of yellow monkeyflower (still affectionately called Mimulus guttatus, Phrymaceae) that carry CMS genes but also nuclear male-fertility restorers (Rf). Carrying both sets of genes means that populations in nature contain only hermaphrodite phenotypes, and female phenotypes are only seen in hybrid crosses between M. guttatus and close relatives (like M.nasutus; see figure to left) that do not carry the appropriate nuclear Rf (see Fishman & Willis 2006 for details on the initial discovery of CMS in M. guttatus!). When CMS always fully restored (i.e., the phenotype is never expressed), it is referred to as 'cryptic CMS.' More on that below! |
Key results of this work to date:
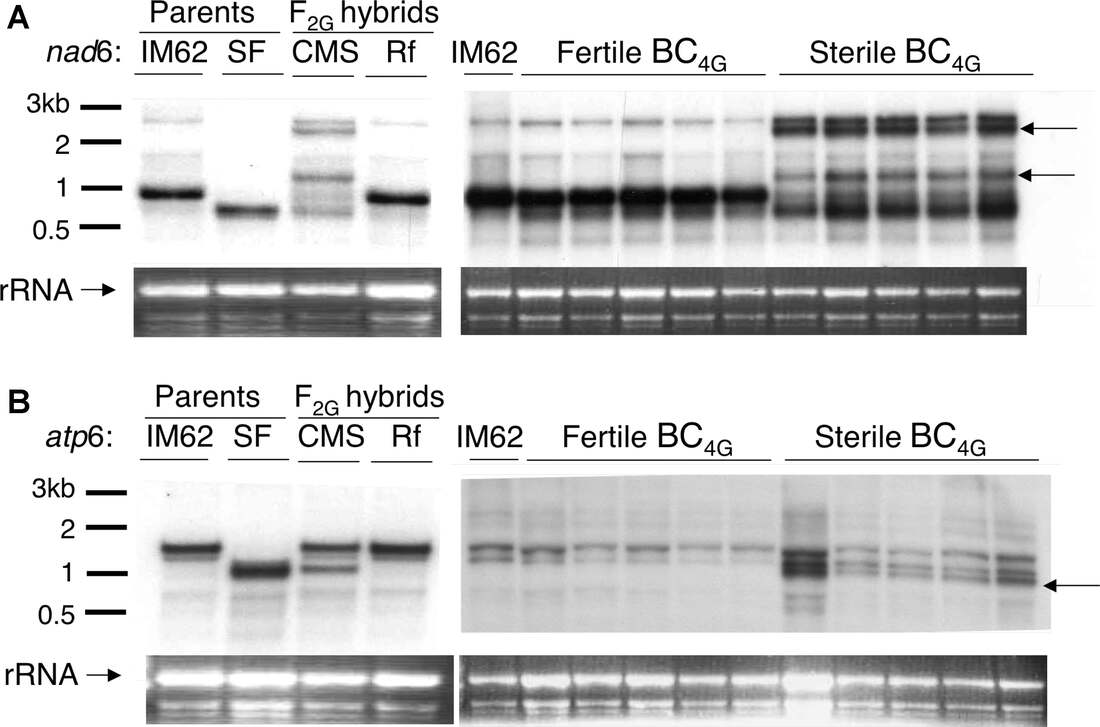
Figure 2. Sterility-associated mitochondrial mRNA in hybrid Mimulus guttatus. (A) Northern blot using labeled nad6 probe identified transcripts abundant in sterile CMS plants (upper arrows) but not their restored fertile full-siblings or the two parental lines (IM62 and
SF). (B) Northern blot using labeled atp6 probe (Case & Willis 2008)
|
|
|
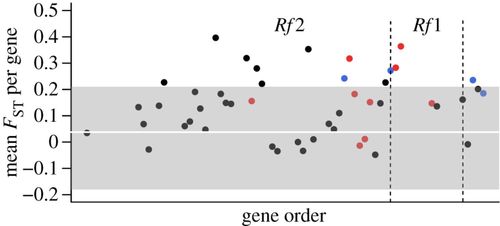
Elevated population-genetic differentiation across the Rf region between M. guttatus populations at Iron Mountain (with IM-CMS) and nearby Cone Peak (without IM-CMS). Mean FST per gene (n = 47) was calculated based on genome re-sequence data from 10 IM lines and 8 CP lines. Genome-wide mean FST is shown by the white line, and 95% confidence limits in grey shading. Red denotes pentatricopeptide repeat genes (=candidate Rf genes). Blue denotes fine-mapped genetic markers used to anchor gene ordering.
|
|